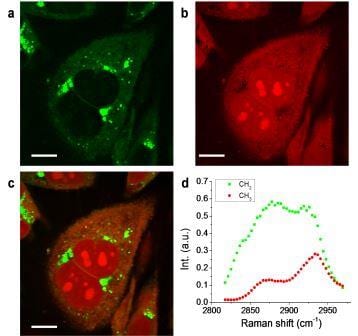
Mapping of the chemical contents and their spatial distribution in a complex biological environment such as cells can be achieved by a new imaging modality: time-lens based hyperspectral stimulated Raman scattering.

Hyperspectral SRS microscopy using a synchronized
time-lens source allows mapping of different cellular
contents.
Label-free imaging of biological samples with endogenous image contrast can be achieved by vibrational spectroscopy. Coherent Raman Scattering (CRS) microscopy, with contrast from coherent anti-Stokes Raman scattering (CARS) or stimulated Raman scattering (SRS), come into consideration as adequate methods. SRS has superseded CARS as a contrast mechanism for microscopy, because it has no image artifacts from non-resonant background and its linear concentration dependence. Yet, it is difficult for single frequency SRS imaging to resolve molecular species that have overlapping Raman bands. Another drawback is the lack of spectral information.
Researchers from Cornell University, Ithaca, and, Purdue University, West Lafayette, now present a new setup that allows for hyperspectral SRS imaging through spectral-transformed excitation.
It is based on a time-lens source synchronized to a femtosecond mode-locked Ti : S laser. The scientists directly modulated the current of the CW laser diode (LD) in the time-lens source with a square wave at a few MHz for SRS imaging, which not only eliminated the need for intensity modulator but also vastly improved the extinction ratio, meaning the optical power ratio between the “on” and “off” state, because below threshold there was no lasing at all. A free-space transmission grating pair with high diffraction efficiency was used to compress the chirped pulse, which produced the modulated output power up to 400 mW. The tunable picosecond pump pulse was generated by pulse shaping of a femtosecond Ti : S laser through intra-pulse spectral scanning.
With a pixel dwell time of 2 ms, the new system was able to image dimethyl sulfoxide (DMSO) in aqueous solution at a concentration down to 28 mM. To demonstrate hyperspectral SRS imaging of biological samples, the team performed SRL imaging of PC3 human prostate cancer cells. Using multivariate curve resolution (MCR) analysis, they decomposed the image into quantitative maps of CH2 largely contributed by lipids, and CH3 largely contributed by protein and nucleotide.
Because the spectral profiles of nucleotide and protein are very similar, the concentration maps of these two components have significant overlaps. The CH2 groups are enriched in lipid droplets and have low concentration inside the nucleus. Meanwhile, the nucleotide showed a distinguishable contribution in nucleoli, indicating their abundance in CH3 groups. Other than MCR, least square fitting can distinguish different components if the Raman spectra of different species in the cell, such as saturated lipid, unsaturated lipid, protein, nucleotide, and water are known.
The researchers expect that the new imaging modality will find wide applications in various fields such as biology, biomedical analysis and diagnosis, and chemical identification.