Many RNAs, including protein-encoding messenger RNA and several types of non-coding RNAs, are transcribed by RNA polymerase II (RNAPII). RNAPII transcription occurs through three highly regulated steps: initiation, when RNAPII is recruited to the DNA as a gene promoter and synthesizes the first few RNA nucleotides; elongation, progression of RNAPII into the gene and extension of the RNA transcript; and termination, release of the RNA transcript and RNAPII from the template DNA. In order to ensure proper transcription at each of these steps, transcription regulatory proteins interact with RNAPII in a dynamic manner. These protein-protein interactions form a highly dynamic and complex network at the site of transcription that allows intricate regulation of gene expression through governing the quality and fate of the transcribed RNA.
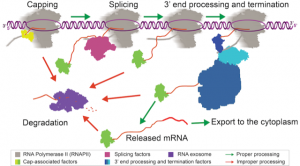
Proper processing and maturation of RNAs is absolutely critical for proper gene expression and protein production. Coupling RNA processing with transcription through protein-protein interactions with RNAPII helps to ensure efficient and proper processing and quality control of different RNA species.
The major processing events that occur co-transcriptionally are 5’ end capping, splicing, and 3’ end processing. If any of these events do not occur properly, the RNA will not be exported from the nucleus and will be restricted to the site of transcription until it can be degraded. RNA transcripts that contain transcription errors or are improperly processed pose a threat to cell health. Therefore, regulation of the processing steps themselves, as well as degradation pathways, act as important quality control strategies to prevent improperly processed RNAs from causing damage to the cell.
A WIREs RNA review highlights recent work that reveals the extent to which RNA processing and quality control are integrated into RNAPII transcription. This review has a particular focus on the balance between 3’ end processing and RNA decay during transcription termination and how that balance plays into RNA quality control.
Kindly contributed by the Authors.











