Viruses cause a wide range of diseases in humans, animals, and plants. Few of these diseases can be effectively treated or prevented by the use of antiviral drugs and vaccines; however, viruses still lead to millions of deaths every year. Viruses are intracellular obligate parasites, which means that they rely on a host cell and its resources to replicate and spread. However, viruses are small and their genetic material only encode for a few proteins. Hence, one of the outstanding questions in modern virology is how a virus conquers the host cell.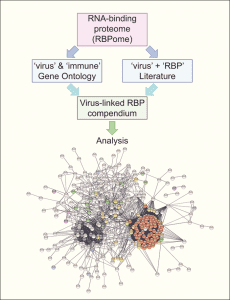
One of the most important tasks of a virus is to replicate its genetic material to form the viral progeny. To achieve this, viruses hijack cellular proteins, many of which are characterized by their capacity to interact with RNA, and are thus called ‘RNA-binding proteins.’ On the other hand, the replication of the viral genetic material is also the ‘Achilles heel’ of viruses, as a specialized group of cellular proteins recognize the different flavors in which the viral genomes are presented, to trigger the antiviral response that sets the infected cells and their neighboring cells into a ‘state of war.’ Many players of host antiviral defenses are RNA-binding proteins. Therefore, cellular RNA-binding proteins emerge as central components of the interactions occurring between a virus and its host cell.
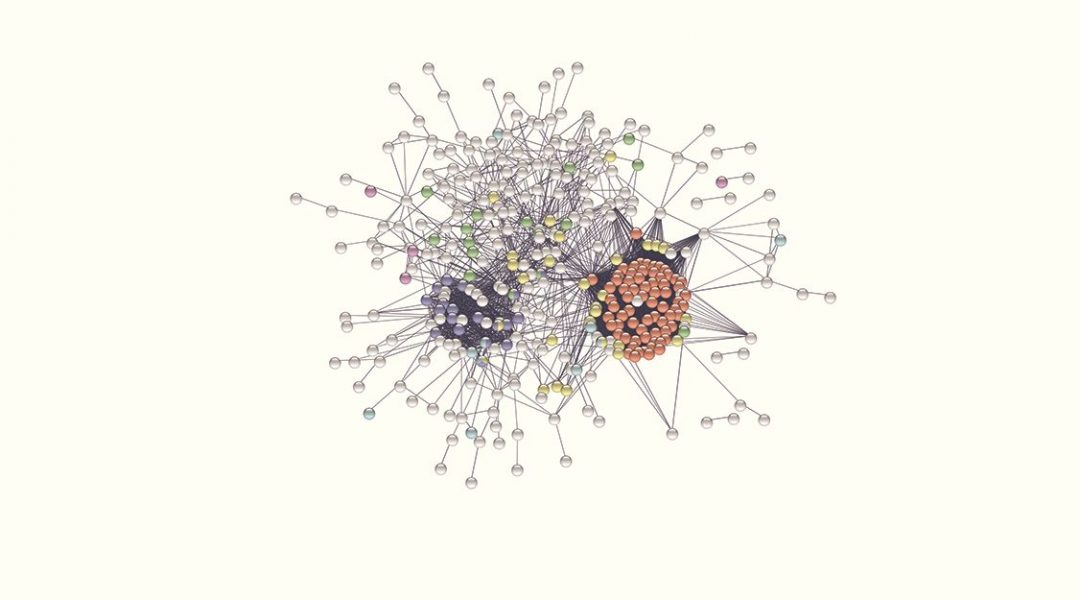
Many efforts have been put towards the identification of the RNA-binding proteins implicated in virus infection. However, the available information on this is fragmented and systematic studies needed to move from ‘individual trees’ to see ‘the forest’ are missing. To fill this gap, a study published in WIREs RNA by Manuel Garcia-Moreno, Aino I. Järvelin and Alfredo Castello took every human protein known to bind RNA and tested the existence of experimental links to virus infection in available databases and scientific literature using computational pipelines.
Applied to the whole repertoire of RNA-binding proteins, they identified near 500 with previous connections to virus infection. While one fifth of these proteins have well-documented modes of RNA binding, the rest were recently described to bind RNA, and thus their mechanisms of action remain poorly understood. Although globally mysterious, the roles of few of these novel RNA-binding proteins in infection have been recently described. In this recent article, the authors discussed these pioneering examples and highlight the need to study the hundreds of cases that remain poorly characterized, as they may represent new targets for next generation antiviral therapies.