Image credit: Kersten Rabe et al.
Antibiotics are becoming increasingly ineffective as drug resistance spreads around the world, making it ever more difficult to treat preventable infections. In addition to the discovery of new antibiotic agents capable of killing resistant bacteria (though according to the WHO “this pipeline is running dry“), understanding how bacteria develop resistance and what leads to the creation of “superbugs” are equally as important in our efforts to stave off this global crisis.
In a recent study published in Small, a team of researchers led by Dr. Kersten Rabe of the Karlsruhe Institute of Technology (KIT) have developed a microfluidic device that allows them to monitor the bacterial evolution of antibiotic-resistant superbugs as a result of exposure to varying levels of antibiotics.
“We wanted to develop and test a microfluidic chip in which cells can be cultured under defined chemical and physical, quasi-static conditions for long periods of time, and to study the influence of these conditions on the occurrence and selection of mutations,” said Rabe.
Research into microfluidic devices and their ability to mimic biological systems has been exploding in recent years, offering inexpensive and robust platforms with striking capabilities such as the ability to mimic organs, model diseases, and test the effects of new drugs, which may be a more effective strategy than animal modeling.
“Microfluidic devices use microscale structures in which liquids and gases are moved under defined flow,” explained Rabe. “This allows, for example, very defined mixing. In addition, reaction parameters such as temperature can be precisely adjusted thanks to the small dimensions and thus small volumes.”
In their paper, the authors say that while miniaturized fluidic chip systems are gaining momentum in the analysis of microbial systems, the ability to generate spatially defined gradients of stressors, such as antibiotics, is still limited, and for the most part, does not allow researchers to carry out long‐term experiments on the bacteria. As a result, they may not be able to discover adaptations that often manifest themselves only after prolonged cultivation.
In microfluidic devices, stressor gradients are usually oriented perpendicular to the system’s fluid flow direction (Figure a, b) and the movement of cells from areas of low stress to high stress must be manipulated, for example, through an arrangement of nutrients: low nutrients at low stress areas and high nutrients at high stress areas to attract cells for screening. Adapted or resistant cells will move into high stress areas, allowing researchers to analyze them. But this is problematic as the adapted cells could potentially never reach the high‐stress areas of the chip where the selection can occur.
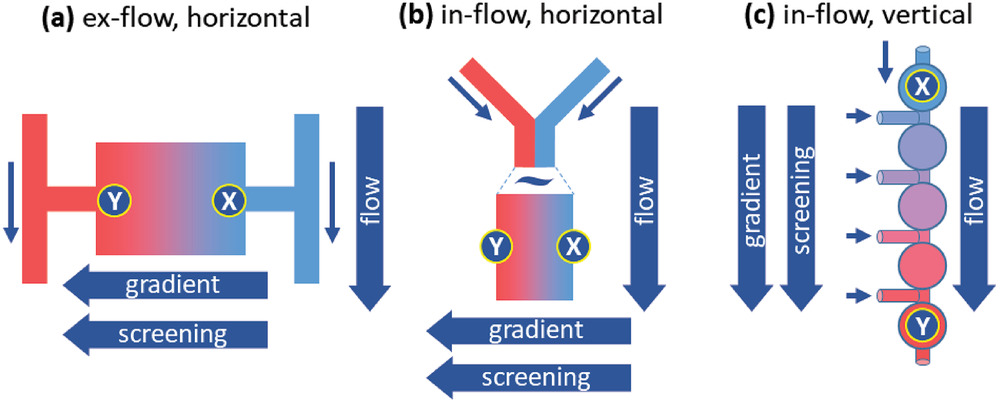
This is what Rabe and his team sought to overcome with a new set up that uses a series of microwells and establishes a stressor gradient along the direction of flow (Figure c). The flow forces carry the cells toward the regions containing high concentrations of the stressor and counteracts a possible evasion of the high stress areas.
“When bacteria are exposed to stress, for example, when they grow in the presence of toxic antibiotics, many cells die,” said Rabe. “However, since the genetic information of the cells is constantly changing as a result of mutations, cells can also be created that are no longer so severely affected by the antibiotics and survive. The chip we have developed makes it possible to select and cultivate such cells preferentially.
“In addition, our chip allows bacteria not only to grow in solution [and are not surface bound], but also to use small wells in the chip to allow bacteria to grow in large communities,” he added. “Such ‘biofilms’ are very common in nature and can play an important role in the development of resistant bacteria.”
The well-based chip allowed the team to better mimic natural microenvironments, such as those found in wastewater systems, exposing the bacteria to defined stressors that promote complex changes in the bacteria, which can now be studied in greater detail.
“Our microfluidic device allows the study of biofilms and biofilm-like cultures with defined stressor concentration gradients and will thus lead to a better understanding of antibiotic resistance mechanisms,” said Rabe. “In this way, previously unknown mutations can be discovered, as was seen [in this work] where we identified specific mutations in the gene involved in the resistance of E. coli to the antibiotic nalidixic acid.”
Rabe also says the chip can be extended beyond microbial resistance mechanisms to the evolution of cancer drug resistance and could even allow scientists to improve industrially relevant microbial strains, such as yeast. In any case, this microfluidic device will allow researchers to generate important data that are needed to better understand the parameters that influence evolution in bacteria, and how these can be overcome to tackle antibiotic resistance.
Reference: Ahmed E. Zoheir, et al., Microfluidic Evolution‐On‐A‐Chip Reveals New Mutations that Cause Antibiotic Resistance, Small (2021). DOI: 10.1002/smll.202007166