At the University of Toronto, Alison McGuigan and her lab focus on the complex and fascinating science of tissue engineering, specifically developing tools to not only mimic the structure of natural tissues, but the processes that lead to their organization.
Her research covers a range of topics from drug resistance in cancer treatment to the use of self-organized 3D tissue cultures called organoids. These are derived from stem cells that have been crafted to replicate an organ or certain arrangements of cells, allowing researchers to study a more realistic model of human tissues.
Beyond her research, McGuigan believes in building communities within academia; environments where students, researchers, and leaders care about one another and drive science through cooperation, support, and collaboration as opposed to a focus on competition.
Our conversation with McGuigan centered around her research and experience as a mother of four leading an academic research group. Her best piece of advice? Practice applying for jobs regularly because you are never going to feel ready when an opportunity comes your way.
Can you tell us about some of your lab’s research in tissue engineering?
Recent work from our lab is centered around mimicking organizational structures that occur during embryonic development. When an embryo develops organs, like the lungs, they go through a series of very specific organizational steps and structures. We wanted to know if these mechanical structures helped define the cell type because you can imagine it is a way of making the whole patterning system more robust. Could we take a big mass of stem cells and put mechanical structural blueprints as well as chemicals to pattern their fate, making them specific lung cells or specific liver cells, for example.
We collaborated with a lung surgeon at the University of Toronto named Thomas Waddell. The idea was to take lungs and measure the bronchial tubes that make up the lung from certain stages of development. We wanted to know if we could control how stem cells differentiate if inserted into different tube sizes corresponding to these different development stages. We were able to make tubes of specific diameter using the same technology that generates microchips. We made plastic surfaces containing holes with dimensions that match those tubes of the lung at different stages of development.
We were able to see that depending on the tube size, you get differences in the way those cells differentiate into particular types of lung cells. That is one aspect of using engineering tools to essentially template the physical organization of cells. You can then drive how they go on to behave in these self-assembly processes where they are differentiating and organizing into 3D structures.
How is your lab able to do this microenvironment engineering with tumor cells?
We have a couple of platforms and there are two criteria that drive what we do.
First, you want to put in the minimum amount of complexity for the problem that you are interested in solving. We used to write in our papers that 2D plastic dishes are not representative of the tumor environment, which contains different cell types, molecular gradients of nutrients and waste, and blood vessels that surround and feed a tumor cell. Tumor cells constantly interact with their microenvironment and [this] potentially can be manipulated to drive tumor cells into particular disease states]. However, one reviewer made a very insightful comment on one of our papers, saying models are not better, it just changes the type of questions you can ask.
Obviously 2D plastic dishes have allowed many cancer drugs to be discovered, so they are very valuable, but they only capture certain kinds of biology.
The second criteria requires that you assemble your structure in a way that facilitates high value data. One example we have is this platform called GLAnCE, which is a culture system we use often with organoids. It simplifies organoid imaging as the hydrogels [used to build them] are fabricated in such a way that they are micro-molded, so their surface is at a very specific thickness and completely flat.
Suddenly, imaging tools work much better because it eliminates a problem presented by a meniscus that forms between the hydrogel and its support surface. It opened up the kinds of data and the quality you can get out your system.
Can you give us a specific example of how you build artificial tumor microenvironments to study diseases?
One of our models of tumor microenvironments is a 3D tool called the TRACER System. To create this, we use a sheet taken from a tea bag — ideal because it’s designed to not fall apart in liquid — and we infiltrate the pores of the teabag with a Jell-O-like material, such as collagen, with the cells inside.
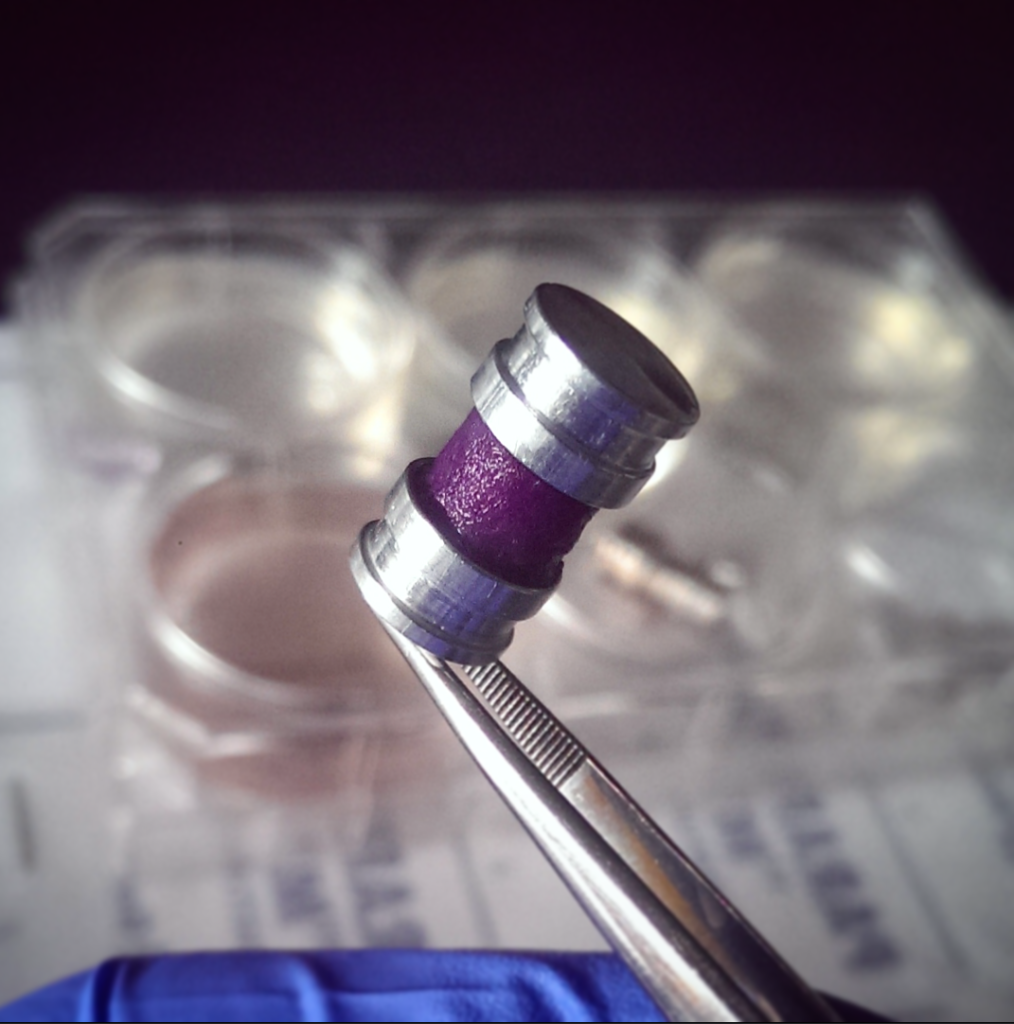
We can then generate this thin “tissue” strip that is about 35 to 40 microns thick. We then roll the strip around an oxygen impermeable core [meaning the oxygen molecules cannot pass through, which effectively mimics a hypoxic or low-oxygen environment typical of tumors], and you end up with something a bit like a Swiss roll, which some people were very confused about in Canada because it is a British cake.
You wrap the cell-Jell-O teabag strip around this core, like a spool of film, and you cap it at either end and then the oxygen and nutrients can only access the stack by diffusing through from the outside towards the center and in no other direction. As such, there is progressive consumption of oxygen in each layer by the cells as the oxygen moves towards the center.
To do analysis, you just unroll the strip and collect cells along its length, tracing back where they were along a 3D gradient. You can rapidly collect cells from different kinds of microenvironments established as a result of this gradient.
Our original system required a lot of cells, but we really wanted to integrate the patient-derived organoid material to capture that more realistic sort of tumor diversity that you get in those populations. We thought it would be easy to just mix them in, but the number of cells you needed to make one model was prohibitive, so we spent a lot of time figuring out how to make the device smaller. The problem is you can only go as small as your ability to roll and handle this strip of tea bag, which is already very small and thin.
A student in my lab, Jose Cadavid, came up with a great idea of filling in the tea bag paper with an oxygen impermeable polymer which leaves regions where you can seed cells. When you roll up the strip, all the regions with cells align and you have a slice of the cake as opposed to the whole circular cake. It is physically the same size as the original model but requires way fewer cells.
How do you assess the behavior of these tumor cells? Is it context specific?
It is context specific. With the TRACER tool, we tried three different organoid lines and observed whether they would develop an oxygen gradient and show oxygen responsive patterns and gene expression, which they all do. Tumor cells in certain regions of the tumor (often where therapies don’t work) are constantly deprived of oxygen and the TRACER tool can mimic that scenario and then investigate how the cells respond at an RNA and eventually protein level.
We have also recently done a single cell RNA-Seq analysis [a technique used to examine the gene expression in only one cell at a given time] through our TRACER platform for one organoid line. This generates a massive amount of information to mine and we can find interesting expression patterns in our data set which seem to be replicating in other organoid models. However, where we have looked more at differences in tumor cell behaviors between patients is in the GLAnCE platform because it has a much higher throughput.
Problems arise when you treat a patient with a drug which kills some proportion of the [cancerous] cells while some remain and then regrow. This recurrence then means those cells may no longer respond to the original drug — you start this “survival of the fittest” cycle.
What our group can do to mimic this is treat our model micro-gels with a drug and then watch them over time. We find situations where we know a certain dose of the drug can kill more cells but the induced changes in the leftover cells actually make them more aggressive and grow back quicker. We have started comparing microenvironments between patients and have found differences in this regrowth behavior.
Many other parameters [of the tumor microenvironment] will also impact how a patient will tolerate a drug [during chemotherapy]and we hope our modeling tools will eventually be able to predict behavior through accurate simulations to be used in a clinical context, not just in the laboratory.
Cell migration leads to the organization of tissues and organs. How and why does your lab investigate cell movement and tissue formation?
Collective migration is fundamental to many processes, and we were studying some of these basic rules but then at some point there was a desire to think about applying our knowledge to different disease situations.
We did some modeling of collective cell migration and determined the rules of behavior in an in vitro system very early on. We also did a lot of theoretical analysis, like tracking how cells move in collective sheets, how they are guided by structural features like topography, and how that guidance could be transmitted between neighboring cells.
In a crowd, if someone is being pulled along and everyone in the crowd is holding hands, you would pull the whole crowd along. At some point, we thought we really wanted to apply this to a more disease-relevant situation, so our attempt at that was to look at the tumor-stroma interface. The stroma is the region found around the tumor and in several cancers, this is primarily fibroblasts. In a number of cancers, you see the breakdown of the tissue structure as the fibroblasts invade into the tumor cells region, and the tumor cells invade out into the stroma compartment.
We have used GLAnCE to study this breakdown of structure in the context of cancer-associated fibroblasts and tumor cells.
Can you share some future research plans?
One paper came out recently that is in a very interesting area I am hoping our lab can expand into. We used the teabag and instead of engineering a tumor, we engineered a sheet of human fat with adipocytes [fat cells] only — there are no other cell types at first. We can then treat this with a high lipid “diet”, simulated in the media, and this induces obese phenotypes. I think this model is going to be an interesting thing to expand on.
Combined with that, we have a method of automation with a robot which could fabricate up to 384 of these engineering fat tissues in parallel. We are going to be able to scale and build these tissues while introducing manipulations to understand how that affects diet-induced phenotypes. We are also starting to explore the molecular pathways that control these processes.
We have some papers coming that introduce immune cells into the cancer system and how we apply these scaled platforms. You can then begin to think about discovering new targets or doing more mechanistic biology.
You are a mom of four. Do you face many challenges as both an academic and a mother?
As a mother, you are forced to prioritize how you want to spend your time. How can you organize your time and identify things that can be as perfect as they can possibly be and other things that can be good enough?
It forces you to get very efficient. I don’t know if my work output changed with each child added, but I think I got more efficient each time. My mind has to adjust between science and then lunches and play dates and this mind switching is probably quite a good thing for your brain. It presumably just changes the way you are seeing these very different worlds and getting comfortable with that switching can be helpful when you then have to collaborate with people that are in very different fields to you.
People are always going to feel guilty that they cannot compete with the metrics of conference attended and hours worked. At the same time, some of the metrics are not relevant to compare anymore because you might go to one conference but be much more targeted about what you are doing at that conference to get the best value.
Travel is really difficult for me, which could make it difficult to get more visibility, but some of the conferences being virtual has probably helped with that, allowing people to attend more meetings without actually having to travel.
I also think building communities on social media is democratizing our ability to get visibility because there is no need for expensive conferences, and people can instead interact with others in the community via Twitter, for example.
Is there something that universities and institutions could do to provide the support system for women as principal investigators?
Many places provide mentorship programs, and I think they require diversity in terms of the mentorship advice because there is not one path to follow. The goal should be to give people confidence to navigate their choices and understand their career options. It would be useful to have some resource that helps you “put out fires” with childcare or elder care, for example.
Being aware of workload in academia is especially important. There is very clear evidence with regards to the need for diversity on award panels, for example, but if you have a smaller fraction of certain demographics in your department, and you need that demographic on every committee, those individuals do much more committee work than other individuals, and that can go unnoticed or is simply not rewarded.
Building a community where people are interested and care about one another as opposed to putting a huge focus on competition is also important. During the COVID-19 pandemic, one professor messaged me asking if I was okay. Honestly, there was nothing they could do, but the fact that they sent the e-mail was amazing.
What is…
A hobby you pursue in your free time?
I would say creative projects, like photo projects or making mini-movies or art.
Do you have a favorite dish?
In Toronto, we are quite spoiled for food. I would say on the different UberEATS options, the whole-chili pakoras are very good. I am vegetarian, so I like a lot of vegan and vegetarian food.
A song you cannot get out of your head at the moment?
I like old school techno house Spotify playlists.
Is there a good book you can recommend?
I recently reread after many years, Tuesdays with Morrie. It is a very sad story full of insight about a man who reconnects with his professor 15 years after being out of university. The professor is dying from Lou Gehrig’s disease, so he wants to give all these lessons to his student, who is writer, before he dies. It is an interesting perspective. I had read it in my mid-20s, and I found it impactful at that time, and it was interesting to read it again and see if I have lived by some of those lessons or not.
A person (famous or not) you would like to have coffee with?
I think it would be interesting to talk to one of these iconic innovators, like Steve Jobs or Elon Musk. Could you have a coherent conversation? Do people like Bill Gates see this nugget of the future? Did they really see the end goal from the beginning? Katalin Karikó is a great modern example of someone I would love to have coffee with. It would also be very interesting to understand some of the female innovators who really drove a vision, like Marie Curie.
Who would play you in a biopic?
One answer from my husband was Drew Barrymore simply because people have told me before that I look like her doppelgänger. The other person I thought of is my oldest daughter who sort of looks like me and is only nine, and right now she wants to be an actress and is very dramatic.
Feature image credit: Neil Ta
Learn more about McGuigan’s work in the Advanced Materials “Women in Materials Science” Virtual Issue — a collection of outstanding work created, steered, and led by female scientists around the world.











