How do cells respond to environmental change? In general, cells must adjust their RNA amounts to appropriate levels with precise timing. To establish such modification in RNA level, synthesis and decay rates of RNAs are tightly regulated. RNA kinetics is therefore a fundamental aspect for gene regulation. Many researchers thus try to understand synthesis (i.e., transcription) and degradation of RNAs.
Recent advance in biotechnologies, such as microarray and RNA-seq, enable us to quantify the RNA amounts of each genes in the cells. By using these technologies, researchers can now study gene-expression at a system-wide level. However, standard RNA-seq can only measure RNA abundance and cannot determine RNA kinetics such as synthesis and decay rate. Use of modified RNA analogs is a prominent solution to quantify RNA kinetics. For example, if we added an uridine analog into the culture medium, cells transcribe newly RNAs with the analog. In this situation, we can distinguish recently transcribed RNAs containing uridine analog from the overall RNA population. By quantifying the amounts of these labelled RNAs, we can determine the synthesis or decay rates of RNAs.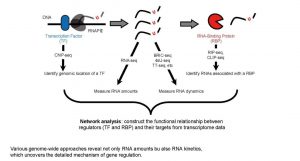
A recent review article in WIREs RNA by Akimsitu and Yamada summarizes the methods that measure the synthesis and decay rate of RNAs. These methods reveal how cells modify mRNA kinetics to establish precise time-course changes of mRNA amounts upon various cellular conditions. Furthermore, the authors discuss the underlying mechanisms of these changes in kinetics, which are regulated by transcription factors and RNA-binding proteins. One important lesson from this review is that both synthesis and degradation of RNAs play a critical role in gene regulation. Considering that many studies only focus on one side of the gene regulation and neglect the effect of the other, combining the methods presented in the review will lead to a comprehensive understanding of gene regulation in cells.
Kindly contributed by the Authors.











