Our increasing ability to perform genetic experiments sometimes leads to bizarre perceptions… Interest in the possibility of humans having mated with chimpanzees at some point, or actually being capable of producing hybrids with chimpanzees, is not merely a perverse public fantasy: several researchers from Russia to China have attempted the crossing, starting in the 1920s and possibly even continuing today. The fact that humans possess one fewer pair of chromosomes than our nearest primate relatives doesn’t seem to dampen enthusiasm for the prospect that a hybrid could be created (true, inequality in chromosome numbers doesn’t absolutely exclude the possibility); and at some point in our ancestry, related species of hominid forebears probably did hybridize, but that doesn’t seem to have occurred after the divergence of Pan (chimpanzees) and Homo (humans).
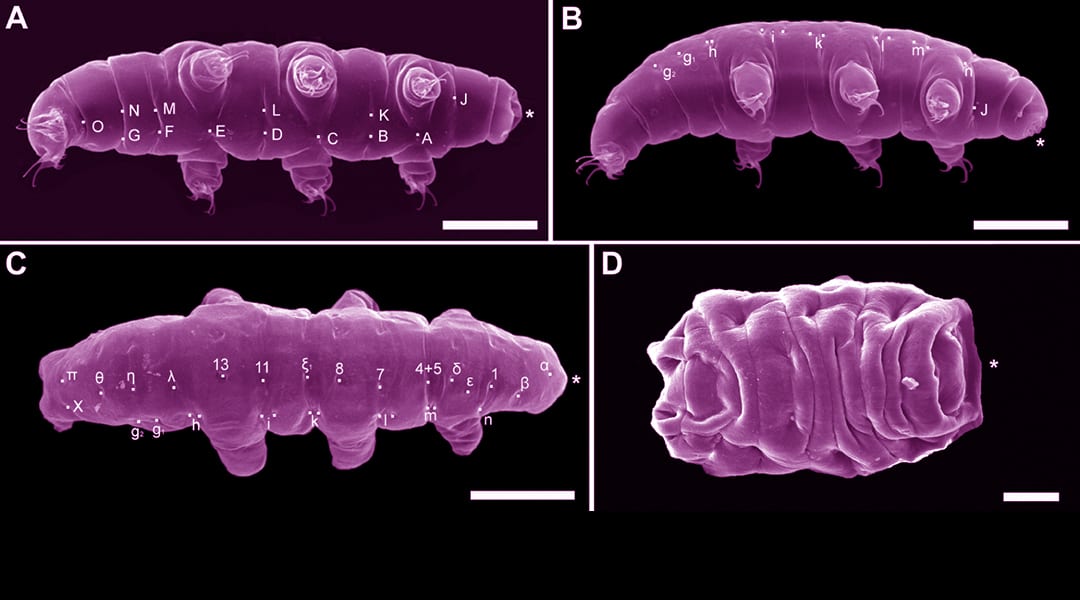
For some reason we are fascinated with the idea that gene pools mix during evolution to produce weird and wonderful things, so if prokaryotes (bacteria and archaea) transfer genes between each other in appreciable amounts, what could be more exciting than showing that organisms much more closely related to us – namely eukaryotes – do the same (most intriguingly, between prokaryotes and eukaryotes)? As objective as we like to think that scientific research is, it seems that there has been a willingness to believe this, and interpret genome analyses accordingly. We were then told, for example, that Tardigrades (micro-animals known colloquially as “water bears” – and, more recently “space bears”(!)) have 17% laterally transferred genes in their genome: well, they were on the rise to ultimate weirdness and wonderfulness, having been discovered (almost) never to have sex (hence not benefiting from the genetic advantages of intra-species exchange), survive decades of desiccation, and – most recently – boast the accolade of being the first animal to survive in the vacuum of space.
It takes a pretty brave soul to diminish the wonder of the Tardigrade by explaining why it can’t possibly be as weird in its genome as it is in other respects; and that is exactly what Bill Martin does in his BioEssays article; but not only does he debunk specific myths of LGT (lateral gene transfer) in eukaryotes, he gives a reasoned scientific argument to explain why we shouldn’t expect much LGT in eukaryotes on the basis of phylogenomics: the nub of the contention is that if LGT is so common in eukaryotes, it must have started a very long time ago, and that would have produced ancestral traits that manifest cumulative effects: basically, such genomic additions, and their subsequent mutations, would have accrued over evolutionary time, and that’s something that we don’t see in phylogenomic analysis. The case is convincingly made in combination with other hard-to-reconcile observations, but the basic point is: “lack of cumulative traits” and plenty of DNA contamination and over-interpretation of sequence analyses. In concert with our increasing skill at genetic analysis, we need increasing scrutiny of how, exactly, it was done, and how the results were interpreted – and a good dose of parsimony…
By Andrew Moore.