 The fundamental question of how life originated is one of the most interesting scientific questions of our time. In particular, the processes by which polymers capable of catalysis and replication (polynucleotides such as RNA and polypeptides or proteins) emerged and propagated on early Earth are still elusive.
The fundamental question of how life originated is one of the most interesting scientific questions of our time. In particular, the processes by which polymers capable of catalysis and replication (polynucleotides such as RNA and polypeptides or proteins) emerged and propagated on early Earth are still elusive.
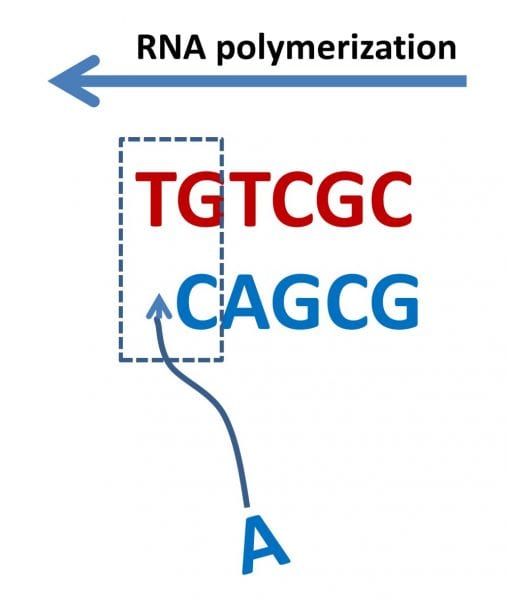
One prevailing theory for the chemical origins of life holds that polynucleotides (RNA, DNA) predate proteins; that there was, in fact, an “RNA world”. In polynucleotide replication, the complement strand, initiated on a primer polynucleotide, is formed by adding nucleotides complementary to a given nucleotide on the template polynucleotide (i.e., adenine to uracil, guanine to cytosine for RNA). Polynucleotide replication, however, is known to be catalyzed by enzymes (proteins) that also suppress errors (mutations) along the chain. In an RNA world, RNA must be capable of replicating through a non-enzymatic mechanism, while retaining the genetic information contained on the template strand. A measure of this capability is the error threshold, defined as the average number of correct nucleotide insertions before an incorrect insertion (error) occurs.
To study this problem, methods of polymer engineering were used by F. Joseph Schork and co-workers at Georgia Institute of Technology to develop a mechanistic model of nonenzymatic polynucleotide replication. When one assumes that multiple incorrect insertions in a row are unlikely, the model returns estimates of the error threshold very much in agreement with the modified Eigen model (based on probability arguments) proposed by Rajamani. In addition, the current model gives average lengths of polynucleotides, categorized by ultimate nucleoside (type and correct/incorrect) as well as the average lengths of interior and terminal sequences of correct and incorrect insertions in a row.