The IL7 receptor (IL7R) and its extra-cellular ligand, the cytokine IL-7 (IL7), are essential drivers of T cell development and homeostasis. Cytokine IL7 is not expressed by T cells but is produced in limited amounts by radio-resistant stromal cells. T cells rely on IL7R signaling and express around 103 copies of the chain IL7Rα (CD127) and 10−105 copies of the γc (CD132) chain (depending on the type of T cell: naive, effector or memory), the two protein sub-units that form the hetero-dimer receptor IL7R. These two facts imply that IL7 signaling at the single T cell level is primarily controlled by IL7 receptor expression, and secondarily by IL7 availability in vivo.
An interesting feature of the IL7 receptor-ligand system in T cells is that the common γ-chain (γc) is shared with a series of other cytokines that include IL-2, IL-4, IL-9, IL15, and IL-21. Since γc expression is presumed to be constitutive and also found in significant amounts in all T cells, many past and current studies of IL7 signaling have focused on regulatory mechanisms of the IL7Rα chain. Notably, the IL7 receptor harbors many unique features that complicate the assessment of IL7R signaling and its downstream effects. For example, IL7 receptor signaling down-regulates expression of its own receptor, so that IL7 signaling leads to suppression of further IL7R signaling. Initiating negative regulatory feedback is quite unusual because expression of other members of the γc receptor family is up-regulated by their cognate cytokine signals. Recent studies have shown that this unique behavior profoundly affects the kinetics and magnitude of IL7 receptor signaling, and that this regulatory mechanism is essential to maintain normal T cell development and homeostasis.
Interrogating how these unique aspects of IL7 receptor signaling are interwoven in the control of T cell development and homeostasis is essential to unravelling the basic mechanisms that regulate T cell-mediated immune responses at both the single cell and population levels. Computational and mathematical models of the dynamical interactions between these many elements (immune molecules and cells) have contributed to our understanding of cytokine receptor signaling, and quantitative approaches and tools have been essential to dissect the contribution of individual nodes in the IL7 signaling pathway.
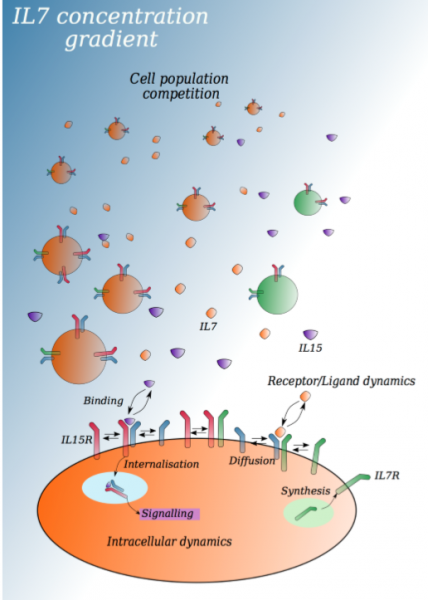
In a WIREs Systems Biology and Medicine review, the authors highlight the current state of basic IL7 receptor biology and focus on the role IL7 and IL7R have on mature CD8+ T cells as drivers of survival and homeostasis. Furthermore, the authors document recent advances in the mathematical and computational modelling of IL7 receptor signaling and their application in furthering our understanding of the dynamics of immune receptor signaling at the molecular, cellular, and population levels.
The review is based on the hypothesis that the development of suitable mathematical models of immune signaling and receptor trafficking will provide answers to some current health-related challenges such as how the expression level (or its copy number) of a given protein in an immune receptor signaling pathway (or network) affect the type and timescale of cellular responses and how ligand concentration or protein competition for binding sites on immune receptors drive different cellular fates by turning on/off different intracellular mechanisms, such as endocytosis, degradation, recycling, or protein synthesis. From a mathematical perspective, the challenge is to develop a quantitative approach to determining how receptor-ligand signaling regulates cellular fate while integrating a wide range of molecular, cellular, and population data. This strategy will help to improve our understanding of the mechanisms that are dysregulated in disease, enabling mathematical models to act as accurate predictors of responses to receptor-targeted therapies and have the potential to aid the design of novel drugs.
Kindly contributed by the Authors.