Alternative splicing is a mechanism that allows eukaryotic, especially mammalian, cells to diversify and control gene expression by creating multiple transcript variants of a single gene. This is done by including or excluding alternative exons. In doing so, the cell modulates protein properties by allowing protein modification, changing protein structure, specifying protein localization, promoting interaction potential, and even altering protein activity. With recent advances in gene editing, DNA and RNA sequencing, and bioinformatics tools, the study of alternative splicing is blooming. Multiple alternative splicing isoforms have been identified and linked to important biological processes, including development and disease.
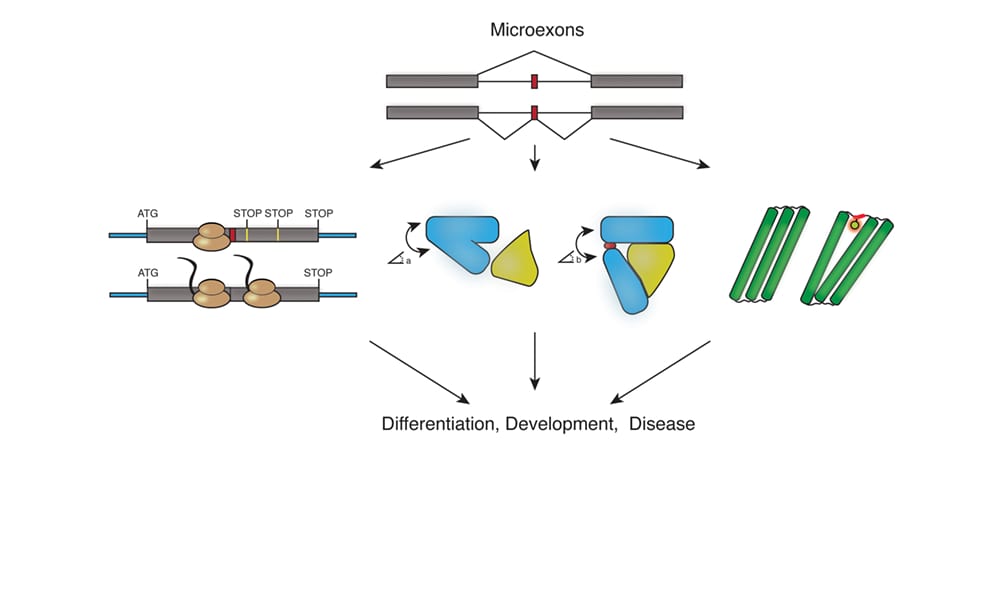
Surprisingly, a unique category of alternative exons has been under studied and considered peculiar outliers. These exons, termed “microexons”, are short 3-27 nucleotide long sequences that encode only a few amino acids in the protein. The short length of microexons makes them difficult to be recognized by the cell splicing machinery, and complicates their identification by scientists. A recent review article in WIREs RNA provides a comprehensive update on the field, describing significant advances made in microexon discovery, including their unique features and the bioinformatics tools that need development to enable their characterization.
Furthermore, the article combines the results of several studies that provide solid evidence for the importance of microexons in the development of various pathologies, particularly their role in the establishment of autism spectrum disorders. Recent works have also identified protein factors responsible for microexon regulation and show an intriguing correlation between microexon splicing and autism in human brains.
Contributed by the Authors.