DNA origami nanostructures were first introduced 10 years ago, and since have become widely used in various, important fields for both applied and fundamental research.
DNA origami can be used as a mask for molecular lithography, as a template for nanoelectronic and plasmonic device fabrication, as a delivery vehicle in molecular machines, and as substrates for single-molecule experiments.
Recently, Adrian Keller and co-workers investigated the stability of DNA origami under strongly denaturing conditions. They wished to elucidate the molecular mechanisms of the attack of denaturants on DNA origami nanostructures, with the hope to provide vital information for future studies on DNA origami-based single molecules in extremely harsh chemical environments.

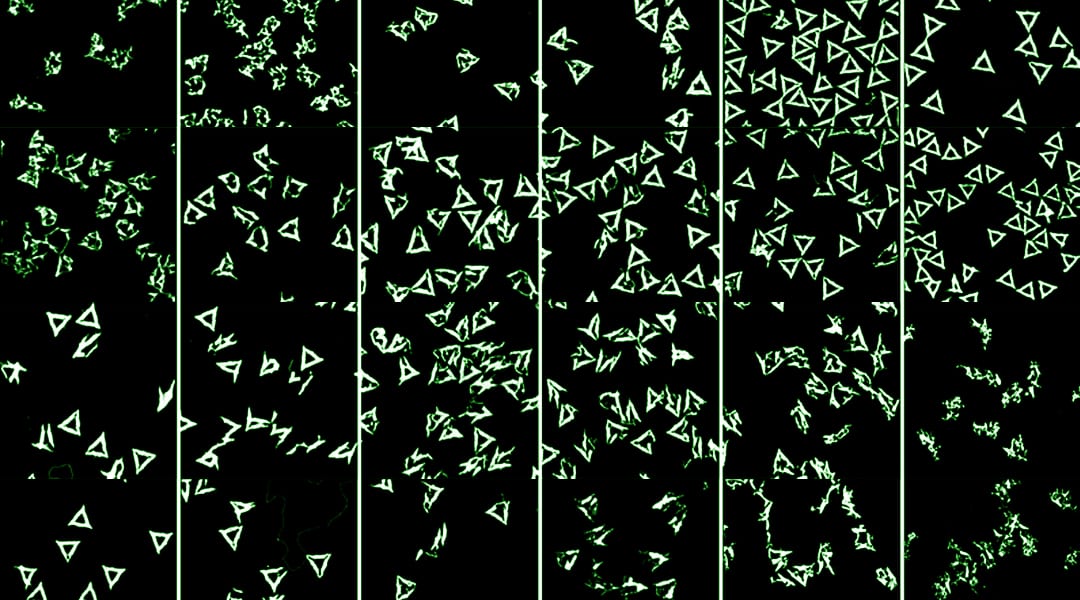
AFM images of DNA origami triangles after 1 h incubation at 42 °C in 7 m urea and different concentrations of NaCl (upper row) and MgCl2 (lower row). Images have a size of 1 × 1 µm2 and height scales are 2 nm.
The effect of Na+ and Mg2+ concentrations on stability was investigated in the presence of two strong denaturants, urea and guanidinium chloride, both commonly employed in protein folding studies. In 4 M guanidinium chloride, denaturation of the DNA origami is promoted by an increased cation concentration.
The authors go on to suggest that cation-induced denaturation by guanidinium chloride deserves to be consider in the future design of single-molecule studies, while potentially being exploitable in denaturation for purification purposes.
If you would like to learn more, please check out the original research article in Small.